Penicillium molds impact the transcriptome and evolution of the cheese bacterium Staphylococcus equorum
Ruby Ye, Christopher Tomo, Neal Chan, Benjamin E Wolfe
mSphere. 2023 May 23;e0004723.
PMID: 37219436 | DOI: 10.1128/msphere.00047-23
Abstract
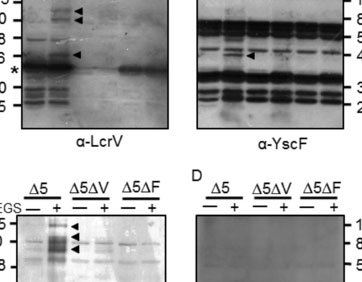
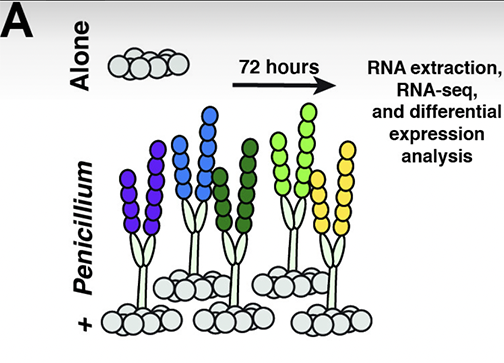
The observation that Penicillium molds can inhibit the growth of Staphylococcus was a catalyst for the antibiotic revolution. Considerable attention has been paid to purified Penicillium metabolites that inhibit bacteria, but little is known about how Penicillium species impact the ecology and evolution of bacteria in multispecies microbial communities. Here, we investigated how four different species of Penicillium can impact global transcription and evolution of a widespread Staphylococcus species (S. equorum) using the cheese rind model microbiome. Through RNA sequencing, we identified a core transcriptional response of S. equorum against all five tested Penicillium strains, including upregulation of thiamine biosynthesis, fatty acid degradation, and amino acid metabolism as well as downregulation of genes involved in the transport of siderophores. In a 12-week evolution experiment where we co-cultured S. equorum with the same Penicillium strains, we observed surprisingly few non-synonymous mutations across S. equorumpopulations evolved with the Penicillium species. A mutation in a putative DHH family phosphoesterase gene only occurred in populations evolved without Penicillium and decreased the fitness of S. equorum when co-cultured with an antagonistic Penicillium strain. Our results highlight the potential for conserved mechanisms of Staphylococcus-Penicillium interactions and demonstrate how fungal biotic environments may constrain the evolution of bacterial species.IMPORTANCEFungi and bacteria are commonly found co-occurring both in natural and synthetic microbiomes, but our understanding of fungal-bacterial interactions is limited to a handful of species. Conserved mechanisms of interactions and evolutionary consequences of fungal-bacterial interactions are largely unknown. Our RNA sequencing and experimental evolution data with Penicillium species and the bacterium S. equorum demonstrate that divergent fungal species can elicit conserved transcriptional and genomic responses in co-occurring bacteria. Penicilliummolds are integral to the discovery of novel antibiotics and production of certain foods. By understanding how Penicillium species affect bacteria, our work can further efforts to design and manage Penicillium-dominated microbial communities in industry and food production.
Keywords: Penicillium; RNA sequencing; Staphylococcus equorum; cheese rind microbiome; evolution; fungal–bacterial interactions.